What is an Ultrametric tree
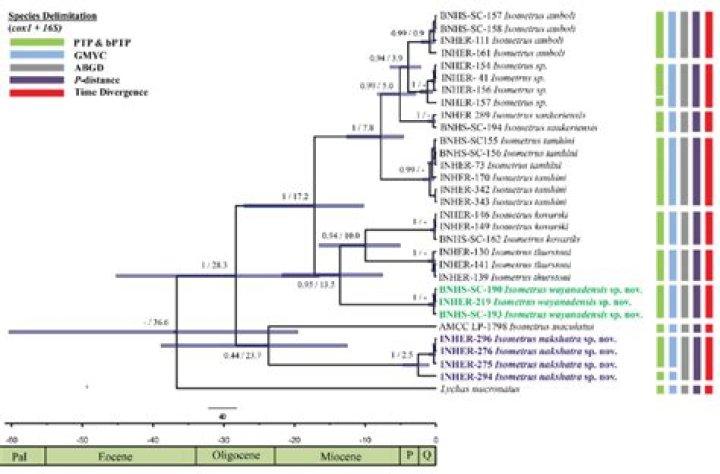
Ultrametric trees (sometimes also called “dendrograms”) are a special kind of additive tree in which the tips of the trees are all equidistant from the root of the tree.
What is an Ultrametric phylogenetic tree?
Ultrametric tree An ultrametric tree or chronogram is a phylogenetic tree where edge lengths represent time (so current taxa are equidistant from the root).
What is the purpose of the phylogenetic tree?
A phylogenetic tree is a visual representation of the relationship between different organisms, showing the path through evolutionary time from a common ancestor to different descendants. Trees can represent relationships ranging from the entire history of life on earth, down to individuals in a population.
Are Ultrametric trees always additive?
– All ultrametric matrices are additive – But, an additive matrix is not necessarily ultrametric.What is an additive tree?
An additive tree is a general way to represent clusters of data in a graph. It is used when your data table is composed of rows and columns that represent the same units; the measure must be a distance or a similarity. A “tree” is a finite, connected graph where any two nodes are connected by one path.
What is ultrametric distance?
More simply, in an ultrametric tree, the distance from the root to the leaves is the same for every leaf. So if all leaf nodes are sampled at the same time and the ultrametric property holds, the tree displaying the distances will have all the leaf nodes at the same level.
What does it mean if two species are sister taxa?
When two lineages stem from the same branch point, they are called sister taxa.
How many internal branches are there in a fully resolved unrooted tree for 5 taxa?
8. Furthermore, we connect the fifth taxon to each branch (5 branches per tree) on each of these 3 trees to yield all 15 possible unrooted trees for the five taxa as shown in Fig. 9.Which type of DNA is useful as a molecular clock?
Mitochondrial DNA is useful as a molecular clock because it displays uniparental inheritance. This is because mitochondrial DNA is only inherited from…
Which is the best definition of a phylogenetic tree?a branching chart that depicts evolutionary relationship among organisms.
Article first time published onWhat is phylogenetic tree in bioinformatics?
A phylogenetic tree (also phylogeny or evolutionary tree) is a branching diagram or a tree showing the evolutionary relationships among various biological species or other entities based upon similarities and differences in their physical or genetic characteristics.
What is a phylogenetic tree quizlet?
Phylogenetic Tree. a diagram designed to reveal evolutionary relationships among DNA or protein sequences by grouping organisms in terms of relative recency (time) of common ancestry. Branch Order.
How can decision tree be improved?
- Add more data. Having more data is always a good idea. …
- Treat missing and Outlier values. …
- Feature Engineering. …
- Feature Selection. …
- Multiple algorithms. …
- Algorithm Tuning. …
- Ensemble methods.
How is a decision tree pruned?
We can prune our decision tree by using information gain in both post-pruning and pre-pruning. In pre-pruning, we check whether information gain at a particular node is greater than minimum gain. In post-pruning, we prune the subtrees with the least information gain until we reach a desired number of leaves.
Is decision tree interpretable?
Decision trees are very interpretable – as long as they are short. The number of terminal nodes increases quickly with depth. The more terminal nodes and the deeper the tree, the more difficult it becomes to understand the decision rules of a tree.
What is the difference between rooted and unrooted phylogenetic tree?
Rooted trees have a single lineage at the base representing a common ancestor that connects all organisms presented in a phylogenetic diagram. … Unrooted trees portray relationships among species, but do not depict their common ancestor.
What is parsimony in evolution?
In general, parsimony is the principle that the simplest explanation that can explain the data is to be preferred. In the analysis of phylogeny, parsimony means that a hypothesis of relationships that requires the smallest number of character changes is most likely to be correct.
Why might a Polytomy exist in a tree?
As suggested, polytomies might be quite common in microbial taxonomy trees since many times evolutionary relationships of interested species cannot be fully resolved to separate descending branches or difference of timeframes between two divergences.
Why do Homoplasious characters arise?
A homoplasy is a shared character between two or more animals that did not arise from a common ancestor. … Often, a homoplasy will occur when two very different groups of animals evolve to do the same thing. This is known as convergent evolution, or convergence. Sometimes, a homoplasy trait is called an analogous trait.
What is the difference between Cladograms and phylogenetic trees?
Cladograms give a hypothetical picture of the actual evolutionary history of the organisms. Phylogenetic trees give an actual representation of the evolutionary history of the organisms. All the branches in a cladogram are of equal length as they do not represent any evolutionary distance between different groups.
Who invented molecular clock?
The concept of a molecular clock was first put forward in 1962 by chemist Linus Pauling and biologist Emile Zuckerkandl, and is based on the observation that genetic mutations, although random, occur at a relatively constant rate.
What is the purpose of molecular clocks?
Instead of measuring seconds, minutes and hours, says Hedges, Penn State professor of biology, the molecular clock measures the number of changes, or mutations, which accumulate in the gene sequences of different species over time.
Why is the use of a molecular clock controversial?
Molecular clocks in general are much more “erratic” than previously thought, and practically useless to keep accurate evolutionary time, the researchers conclude. They attribute this to the vagaries of natural selection, which may at times constrain specific genetic mutations in certain lineages.
What is meant by unrooted tree?
Definitions. A free tree or unrooted tree is a connected undirected graph with no cycles. The vertices with one neighbor are the leaves of the tree, and the remaining vertices are the internal nodes of the tree. … An unrooted binary tree is a free tree in which all internal nodes have degree exactly three.
How do you know if a tree is rooted and unrooted?
# Leaves (n)# Unrooted Trees# Rooted Trees43155151056105945810,395135,135
How do you read an unrooted phylogenetic tree?
Unrooted trees don’t show a common ancestor but do show relationships among species. In a rooted tree, the branching indicates evolutionary relationships (Figure 2). The point where a split occurs, called a branch point, represents where a single lineage evolved into a distinct new one.
What is phylogenetic tree in biology?
Phylogenies are a fundamental tool for organizing our knowledge of the biological diversity we observe on our planet. … A phylogenetic tree, also known as a phylogeny, is a diagram that depicts the lines of evolutionary descent of different species, organisms, or genes from a common ancestor.
How are phylogenetic trees constructed?
Phylogenetic trees are constructed using various data derived from studies on homologous traits, analagous traits, and molecular evidence that can be used to establish relationships using polymeric molecules ( DNA, RNA, and proteins ).
What are phylogenetic species?
oxford. views 1,428,169 updated Jun 11 2018. phylogenetic species concept (PSC) The concept of a species as an irreducible group whose members are descended from a common ancestor and who all possess a combination of certain defining, or derived, traits (see apomorphy).
What is branch in phylogenetic tree?
A branch is called an edge, and represents the time estimate of the evolutionary relationships among the taxonomic units. One branch can connect only two nodes. In a phylogenetic tree, the terminal nodes represent the operational taxonomic units (OTUs) or leaves.
How do you make a simple phylogenetic tree?
Building a phylogenetic tree requires four distinct steps: (Step 1) identify and acquire a set of homologous DNA or protein sequences, (Step 2) align those sequences, (Step 3) estimate a tree from the aligned sequences, and (Step 4) present that tree in such a way as to clearly convey the relevant information to others …